Large-scale scATAC-seq analysis with REMO and BPCells
Compiled: March 23, 2026
Source:vignettes/fetal_remo.Rmd
fetal_remo.RmdView data download code
In this tutorial we will demonstrate how large-scale scATAC-seq datasets from a variety of organs can be efficiently analyzed using Signac with a small memory footprint. This vignette analzying >620,000 cells was run on a Macbook Pro laptop with 16 Gb RAM.
First load in Signac, Seurat, and some other packages we will be using for analyzing human data.
## Warning: package 'Seurat' was built under R version 4.5.2## Warning: package 'sp' was built under R version 4.5.2
library(REMO.v1.GRCh38)
library(BPCells)
library(GenomicRanges)## Warning: package 'GenomicRanges' was built under R version 4.5.2Pre-processing workflow
When pre-processing chromatin data, Signac uses the ATAC fragment file as the main input data. This represents a full list of all unique fragments across all single cells. It is stored on-disk for memory efficiency and scalability. More information about the fragment file can be found on the 10x Genomics website or on the sinto website.
For improved scalability we will use our recently-released REMO features for the human genome, and store these features on-disk using BPCells.
Here we have pre-computed the REMO count matrix using our fragtk Rust software for fast and
memory-efficient matrix quantification, and start by converting this
matrix file to a BPCells matrix:
View fragtk command
# import count matrix
counts <- import_matrix_market_10x(mtx_dir = "remo")
# Write the matrix to a directory in BPCells format
outpath <- paste0(normalizePath(getwd()), "/../bpcells_counts")
write_matrix_dir(mat = counts, dir = outpath)Load on-disk BPCells matrix
remo_matrix <- open_matrix_dir(dir = "../bpcells_counts")
class(remo_matrix)## [1] "MatrixDir"
## attr(,"package")
## [1] "BPCells"Load gene annotations for hg38
library(AnnotationHub)
ah <- AnnotationHub()
ensdb_v98 <- ah[["AH75011"]]
annotations <- GetGRangesFromEnsDb(ensdb = ensdb_v98)
# change to UCSC style since the data was mapped to hg38
seqlevelsStyle(annotations) <- "UCSC"Create Seurat object
chrom_assay <- CreateChromatinAssay5(
counts = remo_matrix,
fragments = "fragments.tsv.gz",
annotation = annotations,
min.cells = 20
)## Computing hash
fetal <- CreateSeuratObject(
counts = chrom_assay,
assay = "REMO",
names.field = 2,
names.delim = "__"
)
fetal## An object of class Seurat
## 338925 features across 626982 samples within 1 assay
## Active assay: REMO (338925 features, 0 variable features)
## 1 layer present: countsDespite containing a large number of cells, we can see that the object requires less that 1 Gb of RAM due to storing the count matrix on disk:
format(object.size(fetal), "Gb")## [1] "0.9 Gb"Quality control
We can compute quality control metrics for the scATAC-seq data using
the ATACqc function. These metrics include:
Nucleosome banding pattern: The histogram of DNA fragment sizes (determined from the paired-end sequencing reads) should exhibit a strong nucleosome banding pattern corresponding to the length of DNA wrapped around a single nucleosome. We calculate this per single cell, and quantify the approximate ratio of mononucleosomal to nucleosome-free fragments (stored as
Nucleosome_signal)Transcriptional start site (TSS) enrichment score: The ENCODE project has defined an ATAC-seq targeting score based on the ratio of fragments centered at the TSS to fragments in TSS-flanking regions (see https://www.encodeproject.org/data-standards/terms/). Poor ATAC-seq experiments typically will have a low TSS enrichment score. Results are stored in metadata under the column name
TSS_enrichment.Total number of fragments: A measure of cellular sequencing depth / complexity. Cells with very few fragments may need to be excluded due to low sequencing depth. Cells with extremely high levels may represent doublets, nuclei clumps, or other artefacts. This is stored as
total_fragments
fetal <- ATACqc(fetal)## Processing /Users/stuartt/Documents/github/vignette_data/zhang/fragments.tsv.gz
DensityScatter(
object = fetal,
x = "total_fragments",
y = "TSS_enrichment",
log_x = TRUE,
log_y = TRUE,
quantiles = TRUE
)## Warning in scale_y_log10(): log-10 transformation introduced infinite values.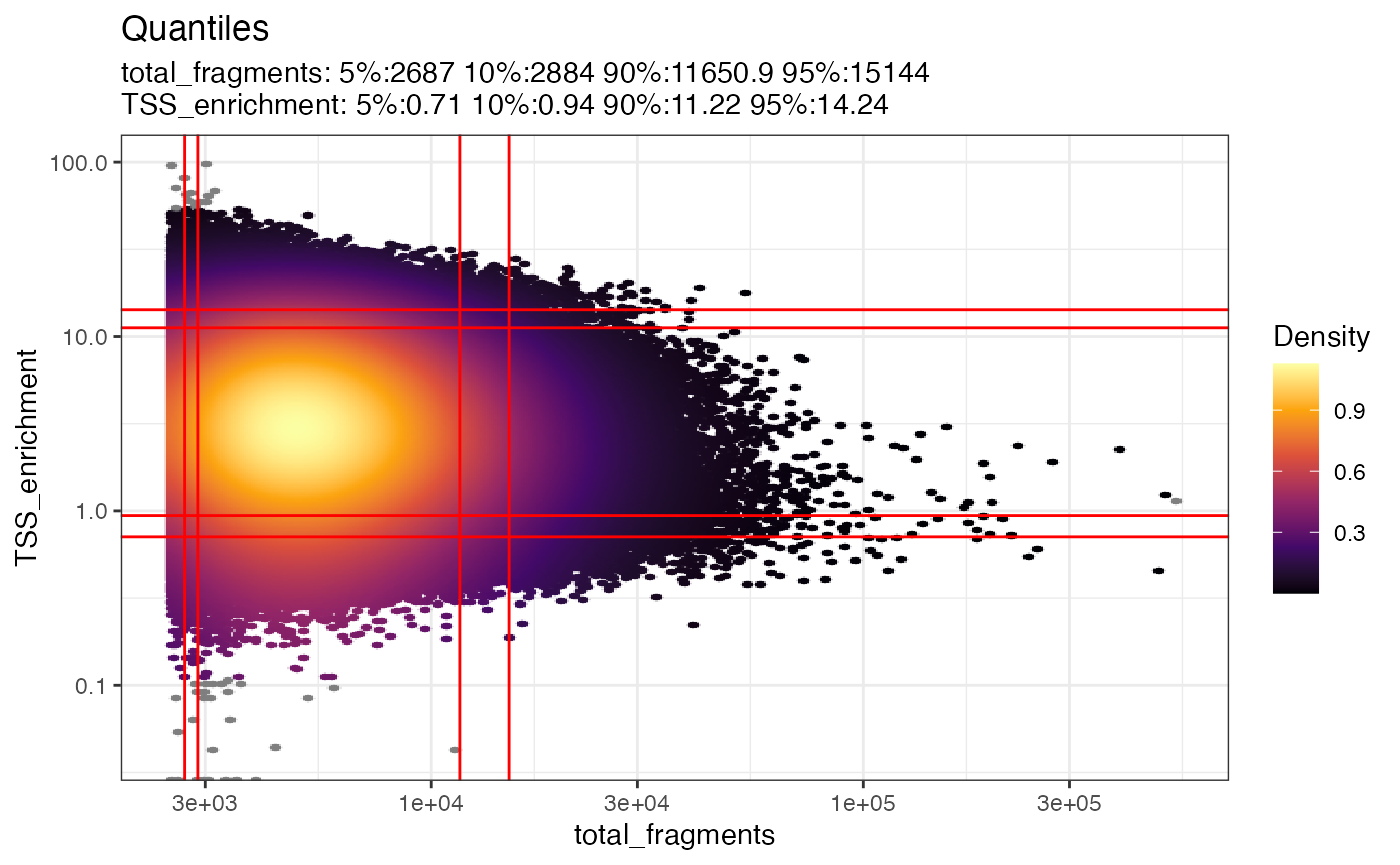
fetal <- subset(fetal, TSS_enrichment > 1)
fetal## An object of class Seurat
## 338925 features across 553575 samples within 1 assay
## Active assay: REMO (338925 features, 0 variable features)
## 1 layer present: countsFeature selection
fetal <- FitMeanVar(fetal)## Finding variable features for layer counts## Retained 338925 features after count filteringNormalization and linear dimensional reduction
fetal <- NormalizeData(fetal)## Normalizing layer: counts
fetal <- RunSVD(fetal, pca = TRUE)## Running PCANon-linear dimension reduction
Now that the cells are embedded in a low-dimensional space we project the cells into a two dimensional space for visualization using UMAP, and color the cells by their original tissue annotation.
fetal <- RunUMAP(object = fetal, dims = 1:50)
# extract original tissue annotations
fetal$tissue <- unlist(lapply(strsplit(as.character(fetal$orig.ident), "_SM-"), `[[`, 1))
DimPlot(fetal, group.by = "tissue", label = TRUE, raster = FALSE, repel = TRUE, pt.size = 0.1) + NoLegend()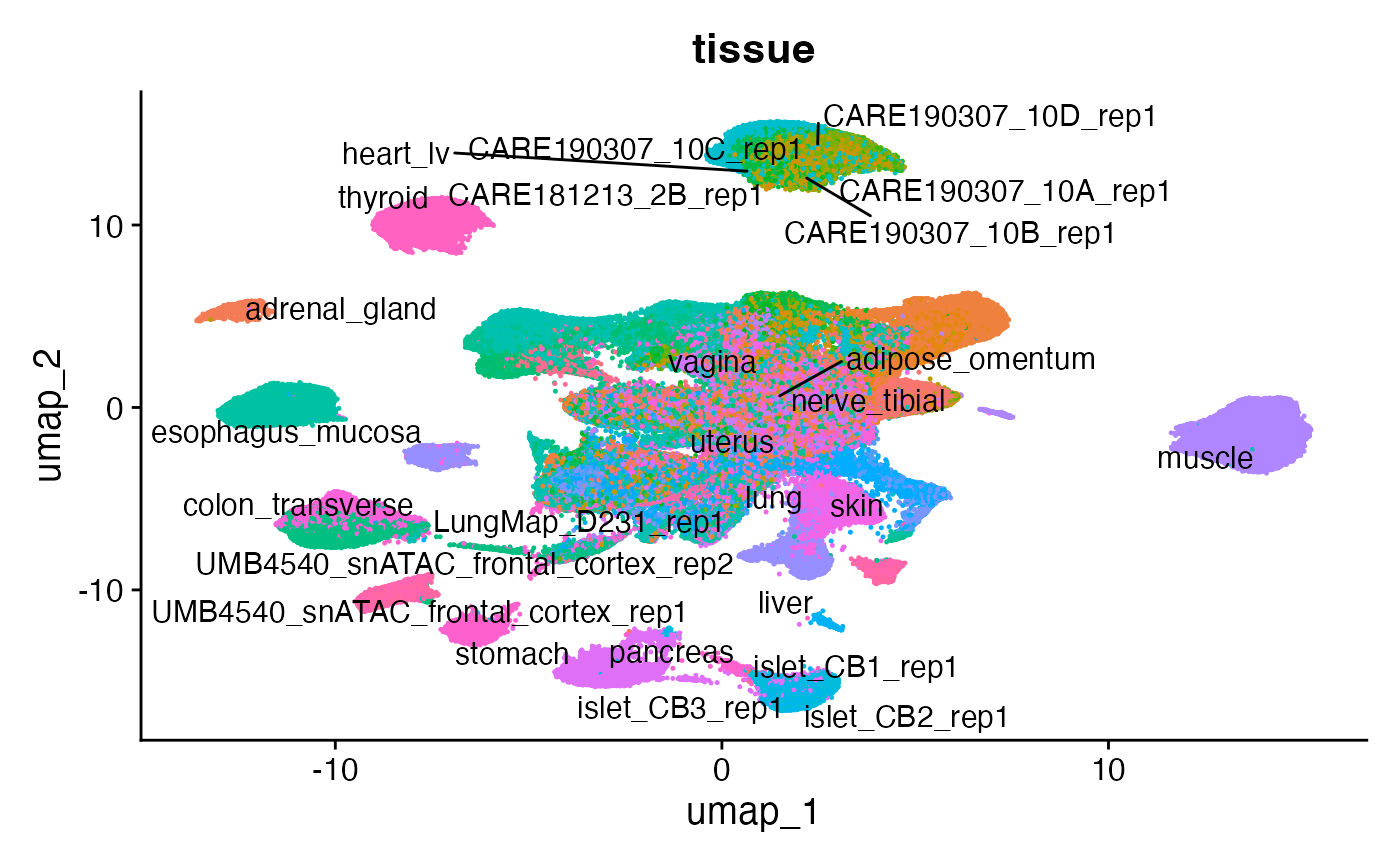
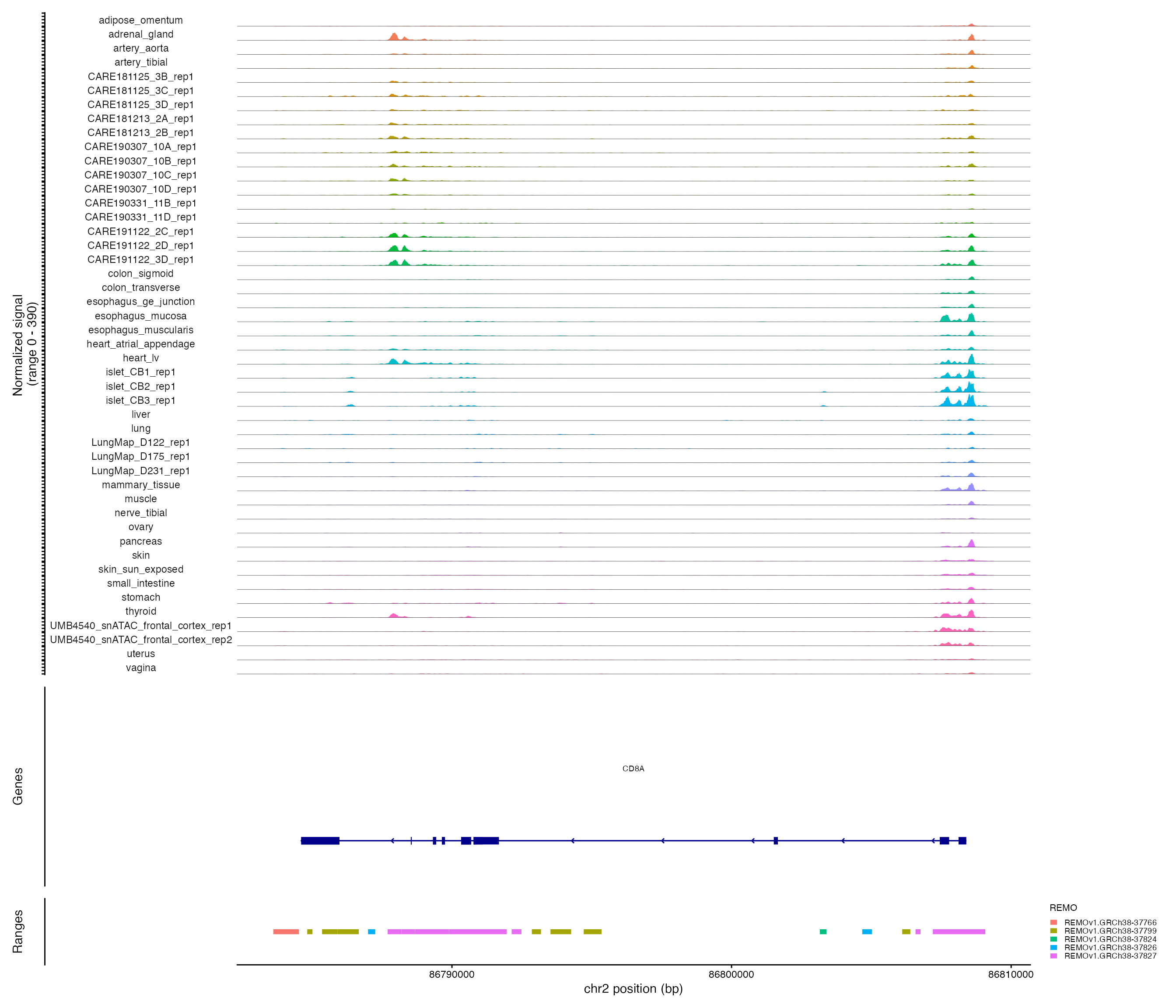
Plotting genomic regions
CoveragePlot(
object = fetal,
group.by = "tissue",
region = "CD8A",
ranges = REMO.v1.GRCh38,
ranges.group.by = "REMO",
extend.upstream = 1000,
extend.downstream = 1000
)
Session Info
## R version 4.5.1 (2025-06-13)
## Platform: aarch64-apple-darwin20
## Running under: macOS Tahoe 26.3.1
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
##
## locale:
## [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
##
## time zone: Australia/Perth
## tzcode source: internal
##
## attached base packages:
## [1] stats4 stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] ensembldb_2.34.0 AnnotationFilter_1.34.0 GenomicFeatures_1.62.0 AnnotationDbi_1.72.0 Biobase_2.70.0 AnnotationHub_4.0.0 BiocFileCache_3.0.0 dbplyr_2.5.2 GenomicRanges_1.62.1 Seqinfo_1.0.0 IRanges_2.44.0 S4Vectors_0.48.0 BiocGenerics_0.56.0 generics_0.1.4 BPCells_0.3.1 REMO.v1.GRCh38_1.0.0 Seurat_5.4.0 SeuratObject_5.3.0.9002 sp_2.2-1 Signac_1.9999.0
##
## loaded via a namespace (and not attached):
## [1] fs_1.6.6 ProtGenerics_1.42.0 matrixStats_1.5.0 spatstat.sparse_3.1-0 bitops_1.0-9 httr_1.4.8 RColorBrewer_1.1-3 InteractionSet_1.38.0 tools_4.5.1 sctransform_0.4.3 backports_1.5.0 R6_2.6.1 lazyeval_0.2.2 uwot_0.2.4 withr_3.0.2 gridExtra_2.3 progressr_0.18.0 cli_3.6.5 textshaping_1.0.4 spatstat.explore_3.7-0 fastDummies_1.7.5
## [22] labeling_0.4.3 sass_0.4.10 S7_0.2.1 spatstat.data_3.1-9 ggridges_0.5.7 pbapply_1.7-4 pkgdown_2.2.0 Rsamtools_2.26.0 systemfonts_1.3.1 foreign_0.8-91 dichromat_2.0-0.1 parallelly_1.46.1 BSgenome_1.78.0 maps_3.4.3 rstudioapi_0.18.0 RSQLite_2.4.6 BiocIO_1.20.0 ica_1.0-3 spatstat.random_3.4-4 dplyr_1.2.0 Matrix_1.7-4
## [43] abind_1.4-8 lifecycle_1.0.5 yaml_2.3.12 SummarizedExperiment_1.40.0 SparseArray_1.10.8 Rtsne_0.17 grid_4.5.1 blob_1.3.0 promises_1.5.0 crayon_1.5.3 miniUI_0.1.2 lattice_0.22-9 cowplot_1.2.0 cigarillo_1.0.0 KEGGREST_1.50.0 pillar_1.11.1 knitr_1.51 rjson_0.2.23 future.apply_1.20.2 codetools_0.2-20 fastmatch_1.1-8
## [64] glue_1.8.0 spatstat.univar_3.1-6 data.table_1.18.2.1 vctrs_0.7.2 png_0.1-8 spam_2.11-3 gtable_0.3.6 cachem_1.1.0 xfun_0.56 S4Arrays_1.10.1 mime_0.13 survival_3.8-6 RcppRoll_0.3.1 fields_17.1 fitdistrplus_1.2-6 ROCR_1.0-12 nlme_3.1-168 bit64_4.6.0-1 filelock_1.0.3 RcppAnnoy_0.0.23 GenomeInfoDb_1.46.2
## [85] bslib_0.10.0 irlba_2.3.7 KernSmooth_2.23-26 otel_0.2.0 rpart_4.1.24 colorspace_2.1-2 DBI_1.3.0 Hmisc_5.2-5 nnet_7.3-20 tidyselect_1.2.1 bit_4.6.0 compiler_4.5.1 curl_7.0.0 httr2_1.2.2 htmlTable_2.4.3 desc_1.4.3 DelayedArray_0.36.0 plotly_4.12.0 stringfish_0.18.0 rtracklayer_1.70.1 checkmate_2.3.4
## [106] scales_1.4.0 lmtest_0.9-40 rappdirs_0.3.4 stringr_1.6.0 digest_0.6.39 goftest_1.2-3 spatstat.utils_3.2-1 rmarkdown_2.30 XVector_0.50.0 htmltools_0.5.9 pkgconfig_2.0.3 base64enc_0.1-6 sparseMatrixStats_1.22.0 MatrixGenerics_1.22.0 fastmap_1.2.0 rlang_1.1.7 htmlwidgets_1.6.4 UCSC.utils_1.6.1 shiny_1.13.0 farver_2.1.2 jquerylib_0.1.4
## [127] zoo_1.8-15 jsonlite_2.0.0 BiocParallel_1.44.0 VariantAnnotation_1.56.0 RCurl_1.98-1.17 magrittr_2.0.4 Formula_1.2-5 dotCall64_1.2 patchwork_1.3.2 Rcpp_1.1.1 reticulate_1.45.0 stringi_1.8.7 MASS_7.3-65 plyr_1.8.9 parallel_4.5.1 listenv_0.10.1 ggrepel_0.9.7 deldir_2.0-4 Biostrings_2.78.0 splines_4.5.1 tensor_1.5.1
## [148] igraph_2.2.2 spatstat.geom_3.7-0 RcppHNSW_0.6.0 reshape2_1.4.5 qs2_0.1.7 BiocVersion_3.22.0 XML_3.99-0.22 evaluate_1.0.5 biovizBase_1.58.0 RcppParallel_5.1.11-1 BiocManager_1.30.27 httpuv_1.6.16 RANN_2.6.2 tidyr_1.3.2 purrr_1.2.1 polyclip_1.10-7 future_1.70.0 scattermore_1.2 ggplot2_4.0.2 xtable_1.8-8 restfulr_0.0.16
## [169] RSpectra_0.16-2 later_1.4.7 viridisLite_0.4.3 ragg_1.5.0 tibble_3.3.1 memoise_2.0.1 GenomicAlignments_1.46.0 cluster_2.1.8.2 globals_0.19.1