Display the genomic ranges in a ChromatinAssay object that fall
in a given genomic region
PeakPlot( object, region, assay = NULL, peaks = NULL, group.by = NULL, color = "dimgrey" )
Arguments
| object | A |
|---|---|
| region | A genomic region to plot |
| assay | Name of assay to use. If NULL, use the default assay. |
| peaks | A GRanges object containing peak coordinates. If NULL, use coordinates stored in the Seurat object. |
| group.by | Name of variable in feature metadata (if using ranges in the Seurat object) or genomic ranges metadata (if using supplied ranges) to color ranges by. If NULL, do not color by any metadata variable. |
| color | Color to use. If |
Value
Returns a ggplot object
Examples
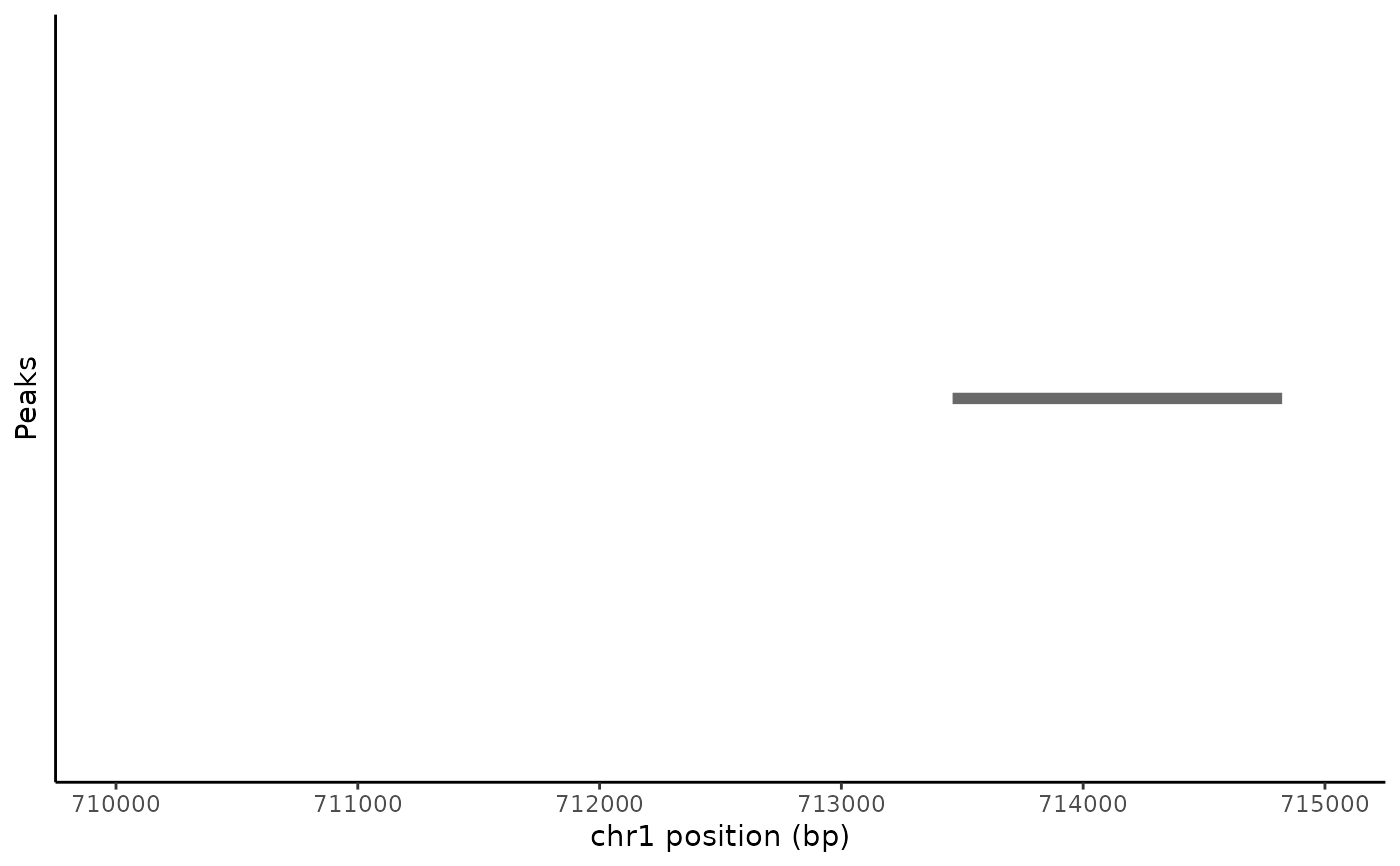
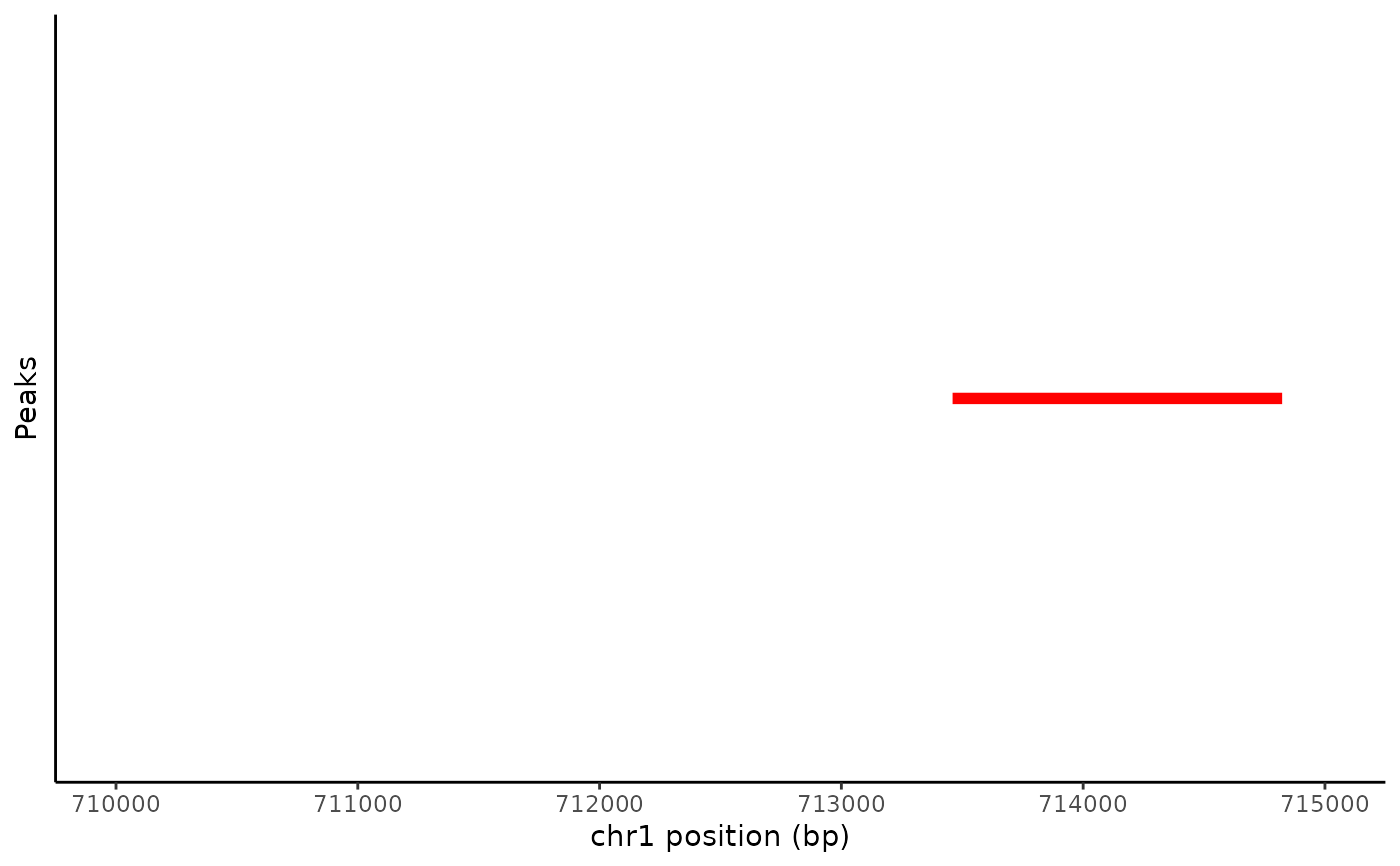
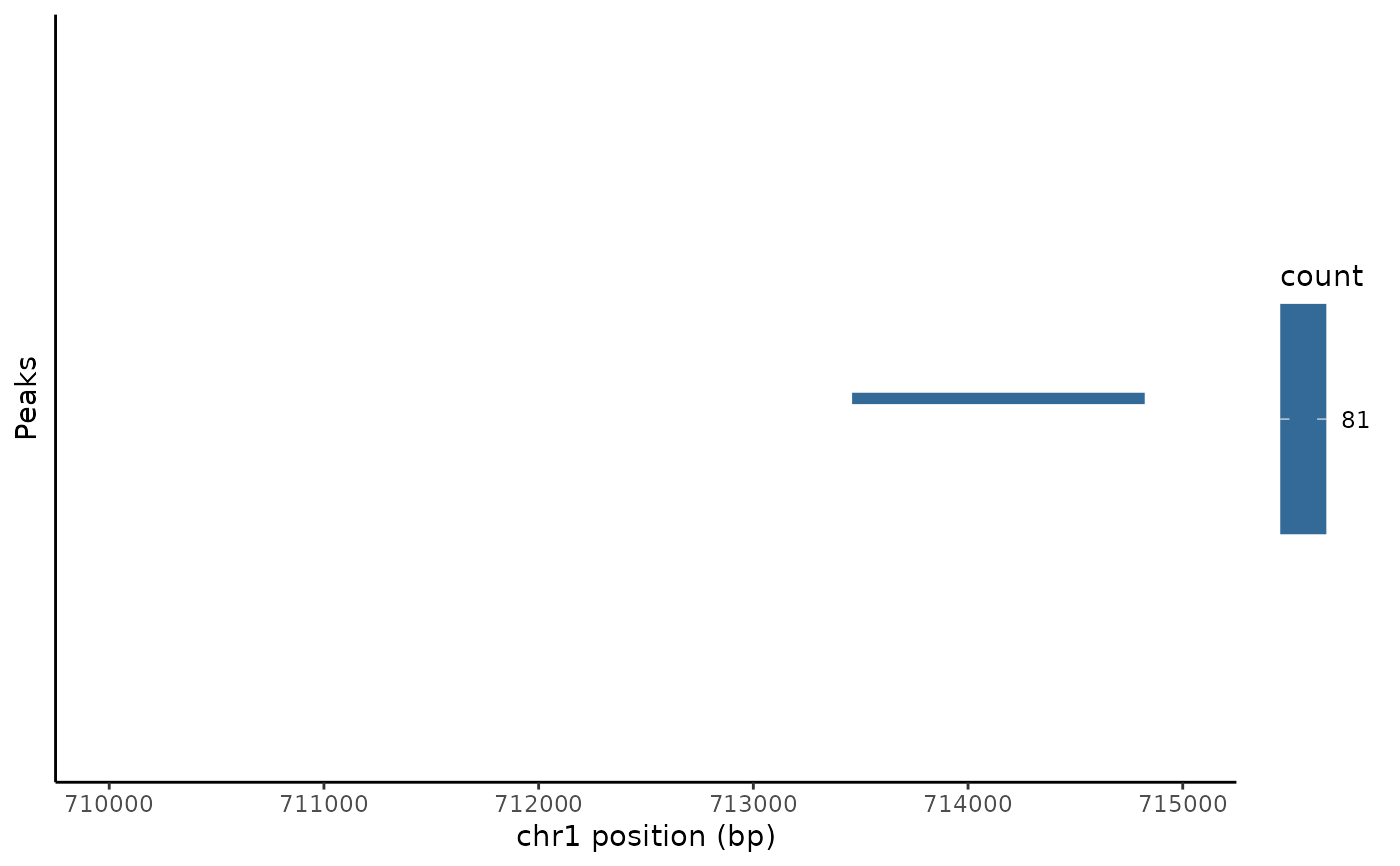
# \donttest{ # plot peaks in assay PeakPlot(atac_small, region = "chr1-710000-715000")# manually set color PeakPlot(atac_small, region = "chr1-710000-715000", color = "red")# color by a variable in the feature metadata PeakPlot(atac_small, region = "chr1-710000-715000", group.by = "count")# }